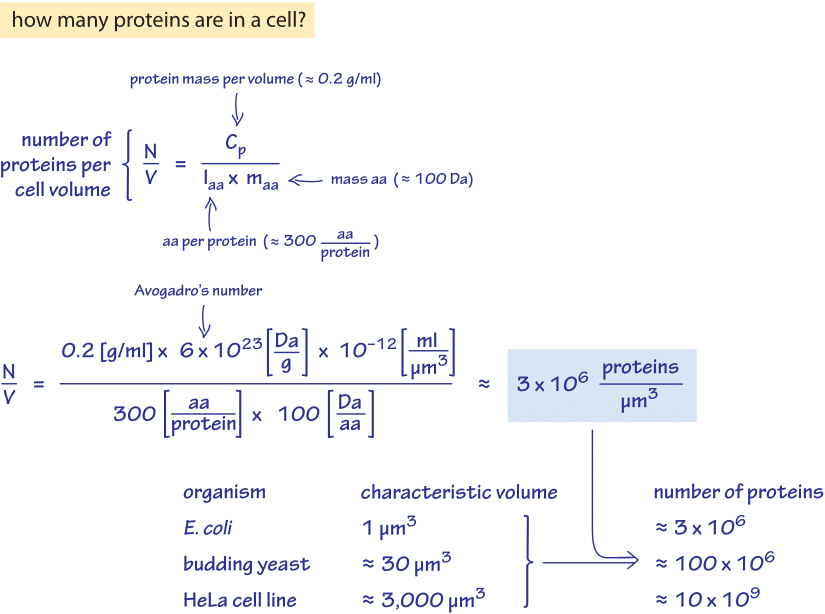
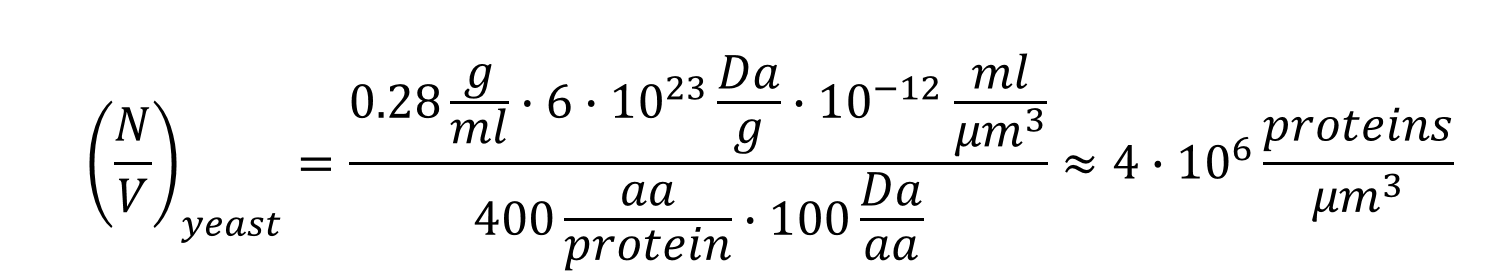What is the total number of protein molecules per cell volume? A call to rethink some published values - Milo - 2013 - BioEssays - Wiley Online Library

PDF) What is the total number of protein molecules per cell volume? A call to rethink some published values
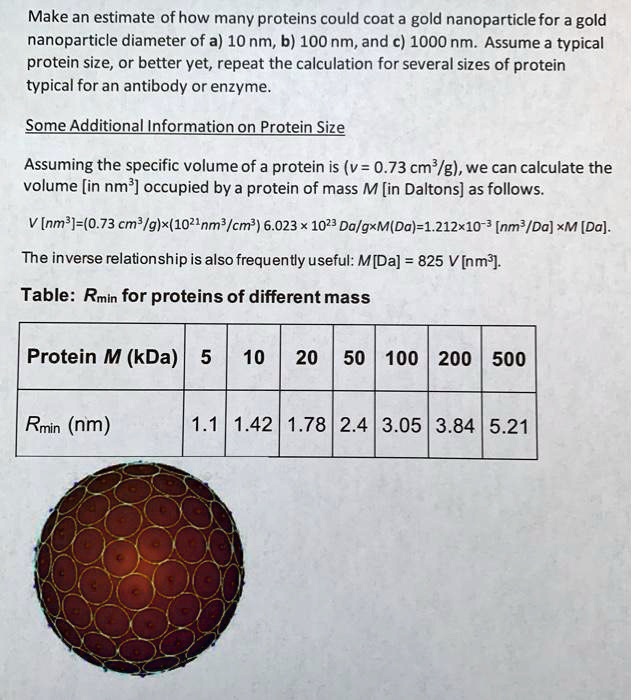

SOLVED: Make an estimate of how many proteins could coat a gold nanoparticle for a gold nanoparticle diameter of a)10 nm,b)100nm,and c)1000 nm.Assume a typical protein size,or better yet,repeat the calculation for

Best practices and benchmarks for intact protein analysis for top-down mass spectrometry | Nature Methods
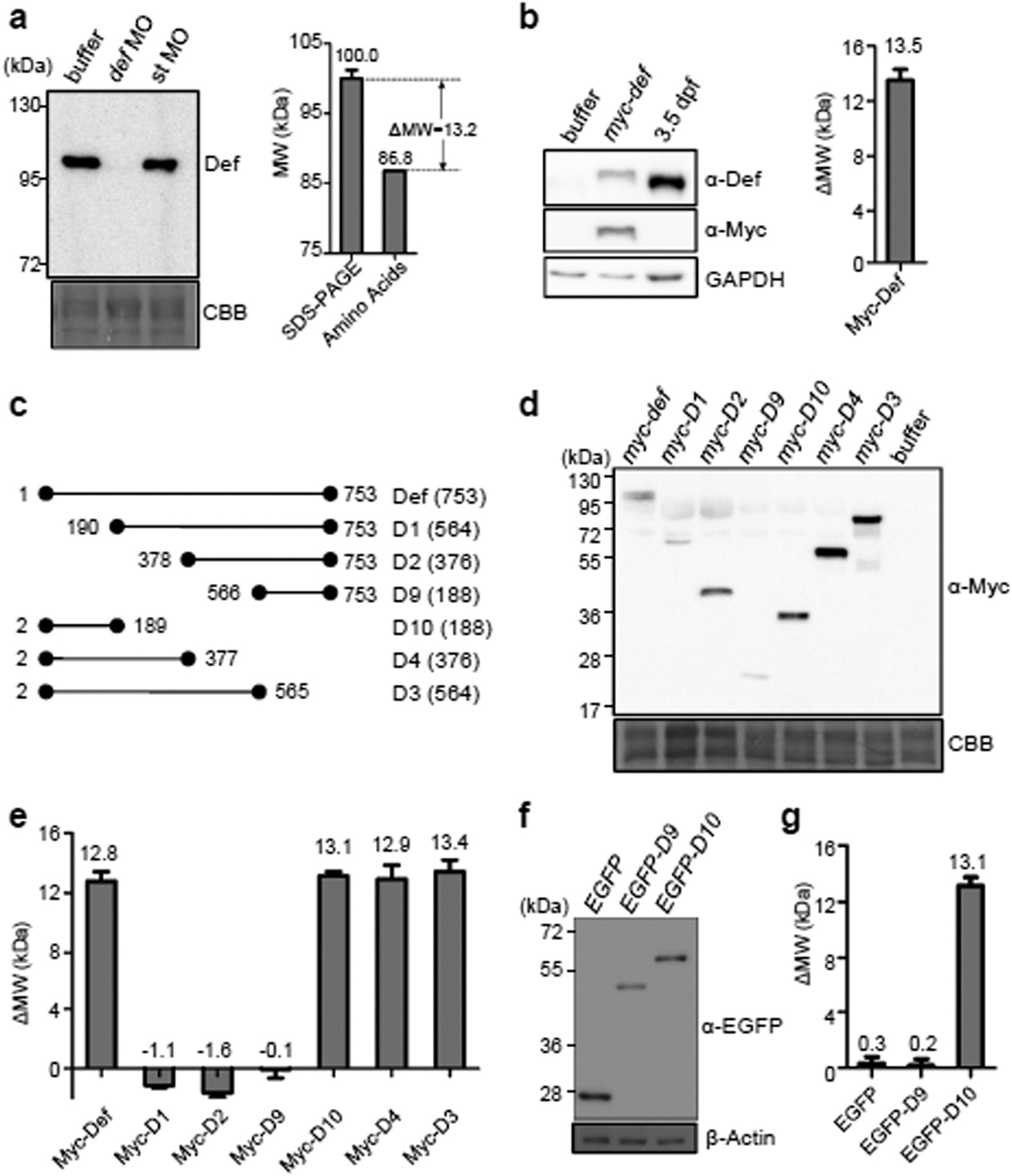
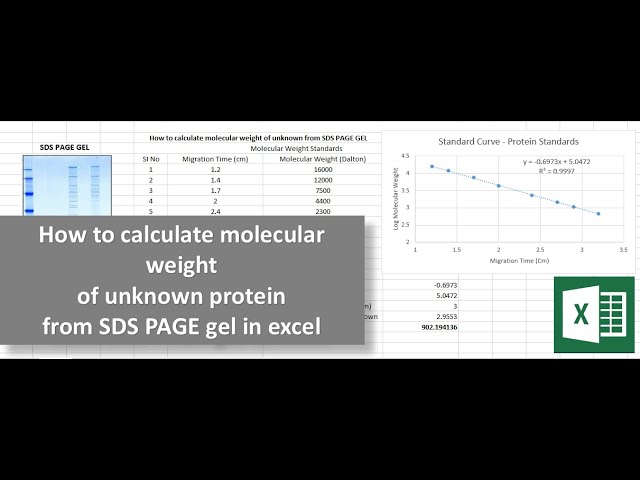
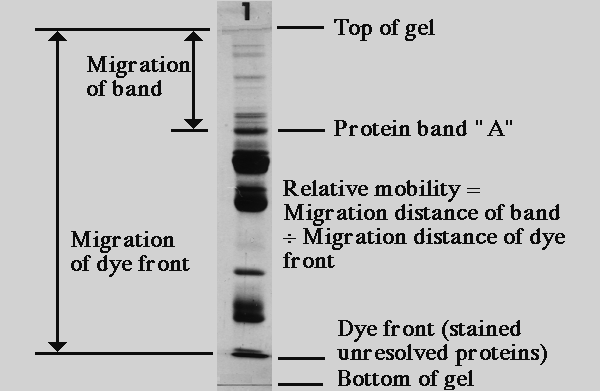
An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide | Scientific Reports

Unification of Protein Abundance Datasets Yields a Quantitative Saccharomyces cerevisiae Proteome - ScienceDirect